High-resolution synchrotron-based X-ray microtomography as a tool to unveil the three-dimensional neuronal architecture of the brain
Matheus de Castro Fonseca, Bruno Henrique Silva Araujo, Carlos Sato Baraldi Dias, Nathaly Lopes Archilha, Dionísio Pedro Amorim Neto, Esper Cavalheiro, Harry Westfahl Jr, Antônio José Roque da Silva, Kleber Gomes Franchini - Brazilian Biosciences National Laboratory (LNBio), Brazilian Center for Research in Energy and Materials (CNPEM), Brazil; Brazilian Synchrotron Light National Laboratory (LNLS), Brazilian Center for Research in Energy and Materials (CNPEM), Brazil; Department of Neurology and Neurosurgery, Federal University of São Paulo (UNIFESP/EPM), Brazil;
![High-resolution synchrotron-based X-ray microtomography as a tool to unveil the three-dimensional neuronal architecture of the brain]()
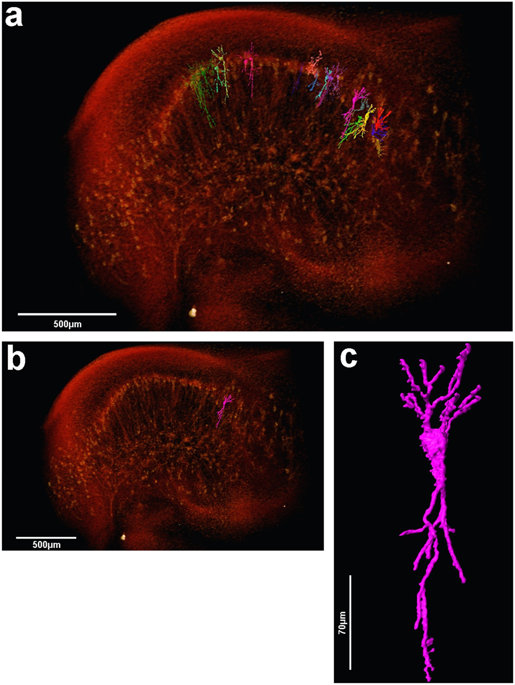
The assessment of neuronal number, spatial organization and connectivity is fundamental for a complete understanding of brain function. However, the evaluation of the three-dimensional (3D) brain cytoarchitecture at cellular resolution persists as a great challenge in the field of neuroscience. In this context, X-ray microtomography has shown to be a valuable non-destructive tool for imaging a broad range of samples, from dense materials to soft biological specimens, arisen as a new method for deciphering the cytoarchitecture and connectivity of the brain. In this work we present a method for imaging whole neurons in the brain, combining synchrotron-based X-ray microtomography with the Golgi-Cox mercury-based impregnation protocol. In contrast to optical 3D techniques, the approach shown here does neither require tissue slicing or clearing, and allows the investigation of several cells within a 3D region of the brain.
How Amira-Avizo Software is used
Image quantification and some specific visualization is only possible from segmented images, i.e., labeled image, where each voxel of the image is correlated to one region of your sample, e.g. cell body, neurons, etc. For that, the 3D reconstructed data obtained at the IMX beamline was post processed using Avizo Fire 9.4, a 3D commercial visualization software. A non-local means filter was applied to all images to remove noise and enable the image segmentation and quantification. Different methodologies of data analysis were applied, depending on the brain region and type of analysis and quantification.