High-resolution structures of HIV-1 Gag cleavage mutants determine structural switch for virus maturation
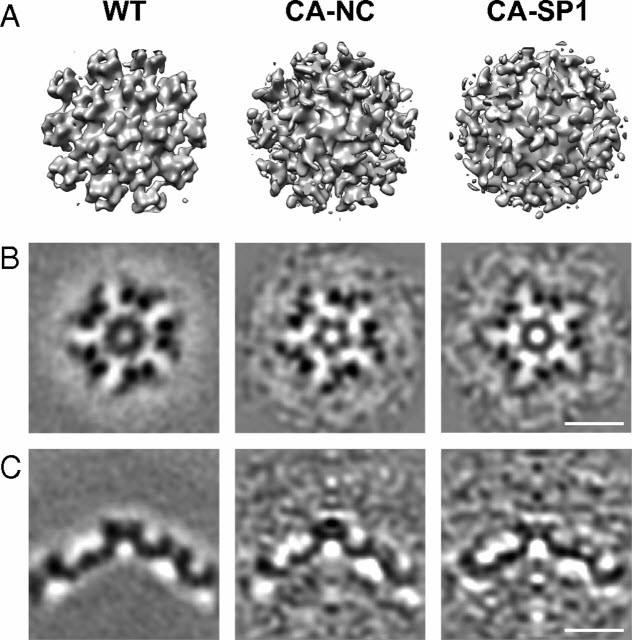
HIV-1 maturation occurs via multiple proteolytic cleavages of the Gag polyprotein, causing rearrangement of the virus particle required for infectivity. (…) How individual cleavages contribute to changes in protein structure and interactions, and how the mature, conical capsid forms, are poorly understood. Here, we employed cryoelectron tomography to determine morphology and high-resolution CA lattice structures for HIV1 derivatives in which Gag cleavage sites are mutated. These analyses prompt us to revise current models for the crucial maturation switch.